Because genotypic information was available for only 257 hybrids, we used the genomic and pedigree relationship matrices to obtain the H matrix for all 415 hybrids. 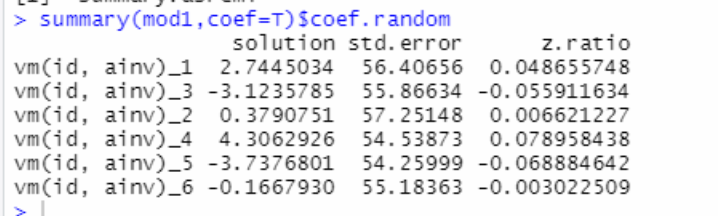
Genotypes of these hybrids were inferred in silico based on their parental inbred lines using single nucleotide polymorphisms (SNPs) markers obtained via genotyping-by-sequencing (GBS). To this end, we used data from 415 maize hybrids evaluated in 4 years of second season field trials for the traits grain yield, number of ears, and grain moisture. Herein, we assess predictive abilities of univariate and multivariate genomic prediction models in a maize breeding program. Therefore, multi-trait multi-environment (MTME) genomic prediction models can leverage these datasets by exploring the correlation between traits and Genotype-by-Environment (G×E) interaction. The dynamics of a commercial breeding program involve the evaluation of several traits simultaneously in a large set of target environments. Especially in maize breeding programs, it emerges as a promising tool for predicting hybrid performance. Genomic selection has become a reality in plant breeding programs with the reduction in genotyping costs. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
